| Issue |
BIO Web Conf.
Volume 27, 2020
International Scientific-Practical Conference “Agriculture and Food Security: Technology, Innovation, Markets, Human Resources” (FIES 2020)
|
|
|---|---|---|
| Article Number | 00066 | |
| Number of page(s) | 5 | |
| DOI | https://doi.org/10.1051/bioconf/20202700066 | |
| Published online | 25 November 2020 | |
Archaea in the microbial community of the reindeer rumen in the Russian Arctic
1
“BIOTROF+” LTD, molecular genetic laboratory, St. Petersburg, 192284, Russia
2
North-west centre of interdisciplinary researches of problems of food maintenance, St. Petersburg, 196608, Russia
* Corresponding author: This email address is being protected from spambots. You need JavaScript enabled to view it.
Abstract
Archaea is the least studied group of the reindeer rumen microbiocenosis. Although the functional load performed by this group of microorganisms in the rumen is large. Methane-forming archaea play a key role in the process of anaerobic decomposition of organic substances, the formation of methane. This study for the first time analyzed the composition of the archaeal part of the microbial community of the reindeer rumen using the T-RFLP method from various regions of the Russian Arctic. As a result, it was found that according to the estimates of the number of archaea by quantitative PCR in the reindeer rumen in the winter-spring period, on average, 108 genomes/g of archaea were observed in individuals of the Yamalo-Nenets Autonomous District, and 109 genomes/in animals from the Nenets Autonomous District Archean. Thus, in the winter-spring period, a lower number of archaea in the rumen was observed in the Yamalo-Nenets Autonomous District. According to the results of the T-RFLP method, 44 to 134 phylotypes were detected in the archaeal community of the reindeer rumen, the Shannon index was 2.02–3.80. The lowest content (up to 11.10 %) of methanogenic archaea of the Methanomicrobia class (including the families Methanosarcinaceae and Methanocorpusculaceae) was revealed in the Nenets Autonomous District, while their presence in individuals of the Yamalo-Nenets Autonomous District reached 36.33 %. Interestingly, in adults of the Yamalo-Nenets and Nenets Autonomous Districts, a significant decrease in the representation of methanogenic archaea of the Methanomicrobia class was noted by 1.38 (P <0.05) and 2.70 times (P <0.01), respectively, compared with young individuals (up to 2 years).
© The Authors, published by EDP Sciences, 2020
 This is an Open Access article distributed under the terms of the Creative Commons Attribution License 4.0, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
This is an Open Access article distributed under the terms of the Creative Commons Attribution License 4.0, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
1 Introduction
Among all ruminants, reindeers are unique animals. They have adaptive devices that help them successfully survive in the Far North [1]. Ruminants are characterized by the presence of a rumen in their digestive system, an organ populated by a large number of microorganisms (bacteria, archaea, fungi, protozoa) that provide fiber digestion. This is a symbiosis that has formed over a long period of co-evolution. According to modern estimates, the diversity of microorganisms in the ruminant rumen reaches several thousand species, less than 100 of which have been studied in detail. Most of them are strictly anaerobic uncultivated species [2].
Thanks to their own enzymatic systems, microorganisms degrade cellulose, lignin and hemicellulose [3], proteins [4], lipids [5], starch [6] into various nutrient compounds. In turn, the host organism creates optimal conditions for the life of microorganisms, i.e. for their growth and reproduction (nutrients, ambient temperature and buffer composition of the environment) [7]. Currently, active research is being conducted on various groups of rumen microorganisms of various ruminants.
One of the groups of the microbial community is methanogenic archaea, which carry out fermentation processes associated with the formation of methane [8, 9]. Methanogenic rumen communities are actively studied by scientists [10, 11]. These communities are microorganisms belonging exclusively to the archaea domain. Methanogens can be divided into free-living and methanogens associated with protozoa and even fungi [12], while the proportion of methanogens in the rumen is about 3 % of all microorganisms. A feature of the simplest rumen is the presence of hydrogenosomes (mitochondria-like organelles), in which hydrogen and carbon dioxide accumulate [13], which makes it possible for methanenes symbionts to actively dwell [14].
Methane-forming archaea play a key role in the process of anaerobic decomposition of organic substances in various ecosystems. These microorganisms are involved in the formation of biogenic methane in such habitats as swamps, rice fields, ruminant rumen and bottom sediments [15, 10].
In general, there are currently few studies on the reindeer rumen microbiome. And quite a few of them relate to such a group of rumen microorganisms as archaea [16]. Therefore, studies of such a group of reindeer rumen microorganisms as Archaea will help expand our understanding of the rumen microbe and a deeper understanding of the adaptive capabilities of these animals. It was shown that methane production increases with an increase in the fraction of indigestible plant feed [17], which is a characteristic feature of the reindeer diet and a high amount of methanogenic archaea in the community can be expected.
This study for the first time analyzed the composition of the archaeal part of the microbial community of the reindeer rumen using the T-RFLP method from various regions of the Russian Arctic.
2 Materials and methods
The object of the study was 38 individuals of reindeer (Rangifer tarandus) of the Nenets breed – calves (4–8 months) and adult individuals (males and females), including young animals (1–2 years). Samples of the contents of the rumen in the winter-spring period were carried out during the slaughter of reindeer: on the territory of the Yamal-Nenets Autonomous District – in the reindeer-breeding brigade of the Yamal Division of the All-Russian Research Institute of Veterinary Entomology and Arachnology and in the Nenets Autonomous District – in the reindeer-breeding brigade of SPK “ERV”. Samples of rumen content to isolate total DNA were frozen immediately after collection and stored at minus 20 °C.
Total DNA from the samples was isolated using the Genomic DNA Purification Kit (Fermentas, Inc. Lithuania), in accordance with the manufacturer’s recommendations.
The amount of DNA extracted was measured using a Qubit® 2.0 fluorometer (Life Technologies, USA). The extracted total DNA was stored at a temperature of minus 20 °C for further analysis.
Evaluation of the bacterial community of the rumen was carried out by T-RFLP (Terminal Restriction Fragment Lengh Polymorphism) and RT-PCR (real-time PCR using 16S rRNA primers (Table 1). Archaeal community taxonomic composition was screened using T-RFLP analysis with using an automatic sequencer CEQ8000 (Beckman Coulter, USA). Sizes and peak areas were calculated in the Fragment Analysis software (Beckman Coulter, USA), as a result of which phylotypes were determined (the error in this study is 1 nucleotide) and their relative share in the community was determined The affiliation of bacteria to the taxonomic group was determined using the database http://mica.ibest.uidaho.edu/trflp.php.
Real-time PCR was performed using a DT-lite amplifier (DNA technology). Using a fluorescent label and specific universal primers for the archaeological community. As a result, real-time PCR using standards took into account the total number of equivalent genomes per gram of archaea.
Mathematical and statistical processing of the results was performed by standard methods of analysis of variance using the software EXCEL XP / 2003 and Past (http://folk.uio.no/ohammer/past/).
To identify a causal relationship between factor and resultant signs, Pearson’s correlation coefficients were calculated, considering them high for absolute values of r greater than 0.6, medium for r more than 0.3 and low for r less than 0.3. Reliability of correlations was determined according to L.S. Kaminsky.
Similarity analysis (ANOSIM) was also used to assess similarity between communities, which is a nonparametric statistical test widely used in the environmental field and uses a ranked matrix of differences.
Characterization of the primers used in the work
3 Results and discussion
According to estimates of the number of archaea by the method of quantitative PCR in the reindeer rumen during the winter-spring period, it was shown that on average 108 genomes/g archaea were observed in individuals of the Yamalo-Nenets Autonomous District, and 109 genomes/g archaea in animals from the Nenets Autonomous District. Thus, in the winter-spring period in the Yamalo-Nenets Autonomous District the lowest number of microorganisms was observed compared to other regions (on average less by 1 order). According to the results of quantitative PCR for vazhenok and choirs, differences by an order of magnitude in the number of archaea between female and male deers in Nenets Autonomous District were noted. At the same time, fewer archaea are specific for males in the Nenets Autonomous District.
In the archaeological community of the reindeer rumen using the T-RFLP method, from 44 to 134 phylotypes were detected depending on the sample; the Shannon index was 2.02–3.80.
When analyzing the archaeal taxonomic affiliation, a significant number of sequences could not be identified up to the phyla level (57.30–96.03 %), which indicates the presence of absolutely unknown microorganisms in the rumen of reindeer, whose nucleotide sequences have no analogues with the described filaments. Unidentified sequences were detected at lower taxonomic levels.
Among the archaeological community of the reindeer rumen, it was possible to distinguish representatives of Phil Crenarheota and Euryarheota, as well as the classes Halobacteria, Methanimocrobia, Methanobacteria, Methanococci and Thermoplasmata. They were present in all studied groups of animals. This is consistent with the data obtained by Sundset et al., Who studied the methanogenic community of the archaeal in rumen of the reindeer of the Svalbard archipelago and found that representatives of the archaea of the classes Methanimocrobia and Methanobacteria were members of this community [16].
In the rumen of cattle, the dominant representative of methanogenic archaea is Methanobrevibacter sp., which can occupy about 65 % of the total methanogenic community [12]. A large number of various methanogens such as Methanobacterium sp., Methanoccccus sp., Methanosphaera sp. carrying out the hydrogenotrophic type of methane biosynthesis, etc. are also present in the rumen of cattle [18, 19].
Assessment of the taxonomic composition of the Archaean community in animals from the Nenets Autonomous District showed the difference between animals from the Yamalo-Nenets Autonomous District. In particular, the lowest content (up to 11.10 %) of methanogenic archaea of the Methanomicrobia class (including the families Methanosarcinaceae and Methanocorpusculaceae) was revealed in the Nenets Autonomous District, while their presence in individuals of the Yamalo-Nenets Autonomous District reached 36.33 %. Also, the archaeological community of the reindeer rumen was characterized by the division of samples into age groups. According to the ANOSIM nonparametric test, the smallest age-related changes in the archaeotic communities of the rumen were revealed for animals from the Nenets Autonomous District.
The results obtained are confirmed by biodiversity indicators for the archaeological community of the rumen during ontogenesis, which varied differently in different regions. For the animals of the Nenets Autonomous District, the environmental indices for young animals and adult animals did not differ significantly, while individuals of the Yamalo-Nenets Autonomous District showed a significant decrease in the Shannon (P <0.05) and Simpson (P <0.05) indices. Judging by the assessment of the taxonomic diversity of archaea, in adult animals of the Yamalo-Nenets Autonomous District and the Nenets Autonomous District, a significant decrease in the representation of methanogenic Archaea of the Methanomicrobia class was noted by 1.38 (P <0.05) and 2.70 times (P <0.01), respectively compared with young individuals (up to 2 years).
According to the results of the ANOSIM similarity analysis, in the Archaean community, the differences between P between young animals and adults were significant in animals of the Yamalo-Nenets Autonomous District (Fig. 4).
Interestingly, in adults of the Yamalo-Nenets and Nenets Autonomous Districts, a significant decrease in the representation of methanogenic archaea of the Methanomicrobia class was noted by 1.38 (P <0.05) and 2.70 times (P <0.01), respectively, compared with young individuals (up to 2 years).
 |
Fig. 1. Average number of microorganisms in the rumen community, lg equivalent genomes/g of content: A – age differences, B – differences by gender |
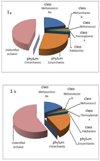 |
Fig. 2. Taxonomic structure of the Archaean community in the reindeer rumen during the winter-spring period from various Arctic regions (T-RFLP analysis): 1) Yamalo-Nenets Autonomous District: a) young growth, b) adult individuals |
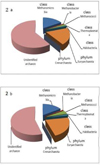 |
Fig. 3. Taxonomic structure of the Archaean community in the reindeer rumen during the winter-spring period from various Arctic regions (T-RFLP analysis): 2) Nenets Autonomous District: a) young growth, b) adult individuals. |
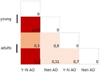 |
Fig. 4. Result of the analysis of the archaeotic community by ANOSIM method |
4 Conclusions
As a result of our study of the community of archaea of the reindeer rumen from various regions of the Russian Arctic, it was found that the number of archaea in the reindeer rumen in the winter-spring period, on average, in individuals of the Yamalo-Nenets Autonomous District was 108 genomes/g of archaea, and in animals from Nenets Autonomous District the figure was 109 genomes/g archaea. Thus, in the winter-spring period, a lower number of archaea in the rumen was observed in the Yamalo-Nenets Autonomous District.
According to the results of the T-RFLP method, 44 to 134 phylotypes were detected in the archaeal community of the reindeer rumen, the Shannon index was 2.02–3.80. The methanogenic archaea of the classes Methanimocrobia, Methanobacteria, and Methanococci made up a large share in the archaean community.
The lowest content (up to 11.10 %) of methanogenic archaea of the Methanomicrobia class (including the families Methanosarcinaceae and Methanocorpusculaceae) was revealed in the Nenets Autonomous District, while their presence in individuals of the Yamalo-Nenets Autonomous District reached 36.33 %. Interestingly, in adults of the Yamalo-Nenets and Nenets Autonomous Districts, a significant decrease in the representation of methanogenic archaea of the Methanomicrobia class was noted by 1.38 (P <0.05) and 2.70 times (P <0.01), respectively, compared with young individuals up to 2 years).
The study was supported by a grant from the Russian Science Foundation No. 17-76-20026 “Microbiocenosis of the rumen Rangifer tarandus of the Arctic regions of Russia as the fundamental basis for obtaining promising biotechnologies for farm animals”.
References
- T.H. Aagnes, W. Sørmo, S.D. Mathiesen, Ruminal microbial digestion in free living, in captive lichen-fed and in starved reindeer (Rangifer tarandus tarandus) in winter, Appl. Env. Microb. 61(2), 583–591 (1995) [CrossRef] [Google Scholar]
- G. Henderson, F. Cox, S. Ganesh, A. Jonker, W. Young, Global Rumen Census Collaborators, Janssen PH., Rumen microbial community composition varies with diet and host, but a core microbiome is found across a wide geographical range, Sci. Rep. 5, 14567 (2015) [CrossRef] [PubMed] [Google Scholar]
- S. Koike, Y. Kobayashi, Fibrolytic Rumen Bacteria: Their Ecology and Functions, Asian-Austral. J. of Animal Sci. 22(1), 131–138 (2009) [CrossRef] [Google Scholar]
- M.A. Brooks, R.M. Harvey, N.F. Johnson, M.S. Kerley, Rumen degradable protein supply affects microbial efficiency in continuous culture and growth in steers, J. Anim. Sci. 90, 4985–4994 (2012) [Google Scholar]
- E.J. Kim, Dietary transformation of lipid in the rumen microbial ecosystem, Asian Austral. J. Anim. Sci. 22, 1341–1350 (2009) [CrossRef] [Google Scholar]
- A. Offner, A. Bach, D. Sauvant, Quantitative review of in situ starch degradation in the rumen, Anim. Feed Sci. Technol. 106, 81–93 (2003) [CrossRef] [Google Scholar]
- E. Rosenberg, G. Sharon, I. Atad, I. ZilberRosenberg, The evolution of animal and plants via symbiosis with microorganisms, Environ. Microbiol. Rep. 2, 500–506 (2011) [CrossRef] [Google Scholar]
- J.M. Brulc, D.A. Antonopoulos, M.E.B. Miller et al., Proc. of the National Acad. of Sci. of the United States of Amer. 106, 1948–1953 (2009) [CrossRef] [Google Scholar]
- R. Wallace, J. Rooke, N. McKain, The rumen microbial metagenome associated with high methane production in cattle BMC, Genomics 16, 839 (2015) [PubMed] [Google Scholar]
- J.I. Velazco, A. Cottle, R.S. Hegarty, Methane emissions and feeding behaviour of feedlot cattle supplemented with nitrate or urea, Animal Product. Sci. 54, 1737–1740 (2014) [CrossRef] [Google Scholar]
- L. Cersosimo, M. Bainbridge, J. Kraft, Influence of periparturient and postpartum diets on rumen methanogen communities in three breeds of primiparous dairy cows, BMC Microbiol. 16, 78 (2016) [CrossRef] [PubMed] [Google Scholar]
- A. Patra, T. Park, M Kim, Rumen methanogens and mitigation of methane emission by antimethanogenic compounds and substances, J. of Animal Sci. and Biotechnol. 8, 13 (2017) [CrossRef] [Google Scholar]
- P.N. Hobson, R.J. Wallace, Microbial ecology and activities in the rumen: Part II. Critical Rev. Microbiol. 9, 253–320 (1982) [CrossRef] [Google Scholar]
- Ch. Newbold, G. de la Fuente, A. Belanch, The Role of Ciliate Protozoa in the Rumen, Frontiers in Microbiol. 6, 1313 (2015) [CrossRef] [Google Scholar]
- K.C. Costa, J.A. Leigh, Metabolic versatility in methanogens, Curr. Opin. Biotechnol. 70, 5 (2014) [Google Scholar]
- M.A Sundset, J.E. Edwards, Y.F Cheng et al., Molecular diversity of the rumen microbiome of Norwegian reindeer on natural summer pasture, Microb. Ecol. 57, 335–348 (2009) [CrossRef] [Google Scholar]
- T. McAllister, K-J. Cheng, E. Okine, G. Mathison, Dietary, environmental and microbiological aspects of methane production in ruminants, Can. J. Anim. Sci. 76, 231–243 (1996) [CrossRef] [Google Scholar]
- M. Auffret, R. Stewart, R. Dewhurst, Identification, comparison, and validation of robust rumen microbial biomarkers for methane emissions using diverse Bos taurus breeds and basal diets, Frontiers in Microbiol. 8, 2642 (2018) [CrossRef] [Google Scholar]
- E.A. Dinsdale., R.E. Edwards, E.D. Frank, J.B. Emerson, P. Wacklin, Gene-centric metagenomics of the fiber-adherent bovine rumen microbiome reveals foragespecific glycoside hydrolases, Proc. Natl. Acad. Sci. USA 106, 1948–1953 (2009) [Google Scholar]
All Tables
All Figures
 |
Fig. 1. Average number of microorganisms in the rumen community, lg equivalent genomes/g of content: A – age differences, B – differences by gender |
| In the text | |
 |
Fig. 2. Taxonomic structure of the Archaean community in the reindeer rumen during the winter-spring period from various Arctic regions (T-RFLP analysis): 1) Yamalo-Nenets Autonomous District: a) young growth, b) adult individuals |
| In the text | |
 |
Fig. 3. Taxonomic structure of the Archaean community in the reindeer rumen during the winter-spring period from various Arctic regions (T-RFLP analysis): 2) Nenets Autonomous District: a) young growth, b) adult individuals. |
| In the text | |
 |
Fig. 4. Result of the analysis of the archaeotic community by ANOSIM method |
| In the text | |
Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.



